$ 0.000 0.01%
Molecular Future (MOF) Rank 2200
| Mkt.Cap | $ 200,028.00 | Volume 24H | 0.00000000MOF |
| Market share | 0% | Total Supply | 200 MMOF |
| Proof type | N/A | Open | $ 0.000002 |
| Low | $ 0.000002 | High | $ 0.000002 |
Molecular Future
TheReaxFF potential[27] can be considered a fully reactive hybrid between bond order potentials and chemistry force fields. First used in theoretical physics, the MD method gained popularity in materials science soon afterwards, and since the 1970s is also common in biochemistry and biophysics. MD is frequently used to refine 3-dimensional structures of proteins and other macromolecules based on experimental constraints from X-ray crystallography or NMR spectroscopy. In physics, MD is used to examine the dynamics of atomic-level phenomena that cannot be observed directly, such as thin film growth and ion-subplantation, and also to examine the physical properties of nanotechnological devices that have not or cannot yet be created. In biophysics and structural biology, the method is frequently applied to study the motions of macromolecules such as proteins and nucleic acids, which can be useful for interpreting the results of certain biophysical experiments and for modeling interactions with other molecules, as in ligand docking.
Molecular Future Technical Analysis & Buy/sell signals

One example of this approach is to use a potential mean force (PMF) which describes how the free energy changes as a particular coordinate is varied. The free energy change described by PMF contains the averaged effects of the solvent. The potential functions representing the non-bonded energy are formulated as a sum over interactions between the particles of the system. The simplest choice, employed in many popular force fields, is the "pair potential", in which the total potential energy can be calculated from the sum of energy contributions between pairs of atoms.

His interests lie in potential blockchain applications which may make a difference to the world. Jason has been closely following technological breakthroughs in the field over the years.
There are a wide variety of semi-empirical potentials, termed tight-binding potentials, which vary according to the atoms being modeled. Semi-empirical potentials make use of the matrix representation from quantum mechanics. However, the values of the matrix elements are found through empirical formulae that estimate the degree of overlap of specific atomic orbitals. The matrix is then diagonalized to determine the occupancy of the different atomic orbitals, and empirical formulae are used once again to determine the energy contributions of the orbitals. Large differences exist between different integrators; some do not have exactly the same highest-order terms as indicated in the flow chart, many also use higher-order time derivatives, and some use both the current and prior time step in variable-time step schemes.

In classical molecular dynamics, the effect of the electrons is approximated as one potential energy surface, usually representing the ground state. A variety of thermostat algorithms are available to add and remove energy from the boundaries of an MD simulation in a more or less realistic way, approximating the canonical ensemble. Popular methods to control temperature include velocity rescaling, the Nosé–Hoover thermostat, Nosé–Hoover chains, the Berendsen thermostat, the Andersen thermostat and Langevin dynamics.
Alternatively, an alignment file can be directly used as input to the program, giving the user control over the alignment method used. In the model building process, the backbone coordinates of the template are copied to the model. If more than one template is used, the SCR from the template with the highest identity is used.
Cloud computing model guarantees scalability advantage because computing resources can be allocated dynamically according to the specific needs of molecular simulation projects. For medium size and little size research centres and/or companies, this issue is fundamental because as soon as the project is done, the cloud resources can be deallocated, avoiding the usage and maintenance of highly costly cluster installed locally. In fact, before, massive simulations were prohibitive due to very long run times (in the order of months) and because of the high cost to establish and maintain powerful computational equipments. In PaaS and SaaS solutions, cloud providers are responsible of maintenance and updating. Ab initio quantum mechanical and chemical methods may be used to calculate the potential energy of a system on the fly, as needed for conformations in a trajectory.

Such united atom approximations have been used in MD simulations of biological membranes. In many-body potentials, the potential energy includes the effects of three or more particles interacting with each other.[30] In simulations with pairwise potentials, global interactions in the system also exist, but they occur only through pairwise terms. In many-body potentials, the potential energy cannot be found by a sum over pairs of atoms, as these interactions are calculated explicitly as a combination of higher-order terms. In the statistical view, the dependency between the variables cannot in general be expressed using only pairwise products of the degrees of freedom. For example, the Tersoff potential,[31] which was originally used to simulate carbon, silicon, and germanium, and has since been used for a wide range of other materials, involves a sum over groups of three atoms, with the angles between the atoms being an important factor in the potential.
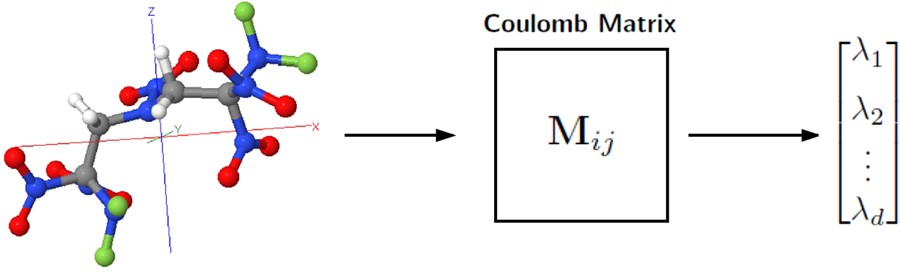
In chemistry and biology this is usually referred to as a force field and in materials physics as an interatomic potential. The simplest form of coarse-graining is the united atom (sometimes called extended atom) and was used in most early MD simulations of proteins, lipids, and nucleic acids. For example, instead of treating all four atoms of a CH3 methyl group explicitly (or all three atoms of CH2 methylene group), one represents the whole group with one pseudo-atom. It must, of course, be properly parameterized so that its van der Waals interactions with other groups have the proper distance-dependence.
Molecular Future Price Index, Historical Chart and Market Cap
When coarse-graining is done at higher levels, the accuracy of the dynamic description may be less reliable. But very coarse-grained models have been used successfully to examine a wide range of questions in structural biology, liquid crystal organization, and polymer glasses.

A lot of important applications of SMD are in the field of drug discovery and biomolecular sciences. SMD was used to investigate the stability of Alzheimer's protofibrils,[53] to study the protein ligand interaction in cyclin-dependent kinase 5[54] and even to show the effect of electric field on thrombin (protein) and aptamer (nucleotide) complex[55] among many other interesting studies. The parameterization of these very coarse-grained models must be done empirically, by matching the behavior of the model to appropriate experimental data or all-atom simulations. Ideally, these parameters should account for both enthalpic and entropic contributions to free energy in an implicit way.
- Molecular Future describes itself as a one-stop digital asset investment service platform jointly invested by Molecular Group, CHINA FORTUNE (00110.HK), Eagle Fund, HBCC Investment, XBTING Foundation, HCASH Foundation, and Collinstar Capital.
- Semi-empirical potentials make use of the matrix representation from quantum mechanics.
- A molecular dynamics simulation requires the definition of a potential function, or a description of the terms by which the particles in the simulation will interact.
- Molecular Future achieves registrations’ volume with 100,000, also manages the fund with 800,000,000 CNY.
Potentials in ab initio methods
Fund management partners are now managing the assets totaling 3.2 billion yuan. Molecular Future achieves registrations’ volume with 100,000, also manages the fund with 800,000,000 CNY.
The issued token by Molecular Future is MOF an ERC 20 Ethereum-based token. MOF is a medium of exchange in Molecular Future ecosystem, acts as a reward payment of the different campaigns running and completed transactions with MOF token. Today Molecular Future price in US dollars is currently 1.20 USD, and if converted to Bitcoin is 0. A total of 11,122 coin's are currently circulating in the Market. Molecular Future prices are currently experiencing a change of -4.55 %, check all cryptocurrency.
Graphics card as a hardware for MD simulations
When there are only 500 atoms, however, the substrate is almost immediately vaporized by the deposition. The temperature of the system in NVE is naturally raised when macromolecules such as proteins undergo exothermic conformational changes and binding.
XBTING enters Molecular Future with debuting the ETH Innovation Fund for new users and others financial products such as current or stable growth cryptocurrency investment plan and hedge fund. They are backed by the resources support of companies listed on Hong Kong, the United States and Australia motherboards. Molecular Future is an innovative financial investment service company registered in the a one-stop digital asset investment service platform. D. C. Rapaport (1996) The Art of Molecular Dynamics Simulation.
The use of non-rectangular periodic boundary conditions, stochastic boundaries and solvent shells can all help reduce the number of solvent molecules required and enable a larger proportion of the computing time to be spent instead on simulating the solute. It is also possible to incorporate the effects of a solvent without needing any explicit solvent molecules present.

The most important advantage of hybrid QM/MM method is the speed. The cost of doing classical molecular dynamics (MM) in the most straightforward case scales O(n2), where n is the number of atoms in the system. This is mainly due to electrostatic interactions term (every particle interacts with every other particle). However, use of cutoff radius, periodic pair-list updates and more recently the variations of the particle-mesh Ewald's (PME) method has reduced this to between O(n) to O(n2). In other words, if a system with twice as many atoms is simulated then it would take between two and four times as much computing power.
In classical molecular dynamics, one potential energy surface (usually the ground state) is represented in the force field. This is a consequence of the Born–Oppenheimer approximation.
In all kinds of molecular dynamics simulations, the simulation box size must be large enough to avoid boundary condition artifacts. Boundary conditions are often treated by choosing fixed values at the edges (which may cause artifacts), or by employing periodic boundary conditions in which one side of the simulation loops back to the opposite side, mimicking a bulk phase (which may cause artifacts too). For simulating molecules in a solvent, a choice should be made between explicit and implicit solvent.
Molecular Future reached its highest price on 6 January, 2020, when it was trading at its all-time high of $ 2.00. You can be signed in via any or all of the methods shown below at the same time. Future developments linking on-line information resources for protein structure to ‘functional datasets’ are illustrated in Fig. 5 of Feedback & CSN access, entry 12, and in Resource J – Search criteria & CSN development.
Molecular Future Project Details
Because of the non-local nature of non-bonded interactions, they involve at least weak interactions between all particles in the system. Its calculation is normally the bottleneck in the speed of MD simulations. To lower the computational cost, force fields employ numerical approximations such as shifted cutoff radii, reaction field algorithms, particle mesh Ewald summation, or the newer particle–particle-particle–mesh (P3M).
In 2016, it achieved a profit of more than 10 million all year round, and in 2017, cooperated with a number of venture capital institutions to jointly set up a special Blockchain investment fund. APP V1.0 of Molecular Future goes online which major in Coins Multiplication and Coins Mortgage with providing a super flow entrance for the organizations and personal investors, also will create a board of “Molecular Exponentiation” for the MOF holders. Li Boji is the recipient of Economics and Psychology in UBC University in Canada, and chairman of XBTING Foundation. He is a continuous entrepreneur for many years and mainly engaged in Blockchain project investment. Since 2013, he began to contact bit currency and has participated in early investment of several star projects.
As the Chief Technology Officer, Jason is focused on the development of the project's technological infrastructure, ensuring the functionality and efficiency of the transaction systems. Additionally, Jason is responsible for the network security of the Molecular Future group. Combined with complete ecological support, start-up companies that hundreds of funds have invested in will gradually march into CTD field through Molecular Future at the appropriate time. Molecular Future Foundation will participate in supporting and jointly entering the investment industry at early stage.
Therefore, these force fields are also called "additive force fields". An example of such a pair potential is the non-bonded Lennard–Jones potential (also termed the 6–12 potential), used for calculating van der Waals forces. Chemistry force fields commonly employ preset bonding arrangements (an exception being ab initio dynamics), and thus are unable to model the process of chemical bond breaking and reactions explicitly.







